| Entry |
Thumbnail |
Name |
Description |
Object |
Legend |
|
hsa00970
|

|
Aminoacyl-tRNA biosynthesis
|
|
C06113 (L-Aspartyl-tRNA(Asn))
C02992 (L-Threonyl-tRNA(Thr))
C02839 (L-Tyrosyl-tRNA(Tyr))
C00078 (L-Tryptophan)
C03512 (L-Tryptophanyl-tRNA(Trp))
C00082 (L-Tyrosine)
C00188 (L-Threonine)
C00123 (L-Leucine)
C02047 ...
|
L-Aspartyl-tRNA(Asn)
6.1.1.5
AMINOACYL-tRNA BIOSYNTHESIS
6.1.1.22
6.1.1.21
6.1.1.20
6.1.1.19
6.1.1.18
6.1.1.16
6.1.1.14
6.1.1.12
6.1.1.11
6.1.1.10
6.1.1.9
6.1.1.7
6.1.1.6
6.1.1.4
6.1.1.3
6.1.1.2
2.1.2 ...
|
|
hsa00980
|

|
Metabolism of xenobiotics by cytochrome P450
|
|
C14039 (1,1-Dichloroethylene)
C14876 (S-(2-Hydroxyethyl)-N-acetyl-L-cysteine)
C14875 (S-(2-Hydroxyethyl)glutathione)
C14877 (S-[2-(N7-Guanyl)ethyl]-N-acetyl-L-cysteine)
C14874 (Glutathione episulfonium ...
|
1,1-Dichloro-ethylene
CYP2E1
CYP2E1
CYP2E1
CYP2E1
CYP2E1
S-(2-Hydroxyethyl)-N-acetyl-L-cysteine
S-(2-Hydroxyethyl)-glutathione
S-[2-(N7-Guanyl)ethyl]-N-acetyl-L-cysteine
Glutathione-episulfonium ion
Thiodiacetic ...
|
|
hsa00982
|

|
Drug metabolism - cytochrome P450
|
|
C16544 (alpha-Hydroxytamoxifen)
D00343 (Ifosfamide (JAN/USP/INN)), C07047 (Ifosfamide)
C16612 (Citalopram aldehyde)
C16648 (2-n-Propyl-4-pentenoic acid)
C16649 (4-Hydroxyvalproic acid)
C16650 (5-Hydroxyvalproic ...
|
α-Hydroxytamoxifen
Ifosfamide (prodrug)
Citalopram-aldehyde
4-Ene-VPA
4-OH-VPA
5-OH-VPA
Oxcarbazepine
Felbamate
4-Hydroxy-5-phenyltetrahydro-1,3-oxazin-2-one
5-Phenyl-1,3-oxazinane-2,4-dione
3-Carbamoyl-2-phenyl-propionic ...
|
|
hsa00983
|

|
Drug metabolism - other enzymes
|
|
D00346 (Isoniazid (JP18/USP/INN)), C07054 (Isoniazid)
C04646 (6-Thioinosine-5'-monophosphate)
D01223 (Capecitabine (JAN/USP/INN)), C12650 (Capecitabine)
C07446 (Isonicotinic acid)
C16614 (6-Methylmercaptopurine)
C16615 ...
|
Isoniazid
6-Thioinosine-5'-monophosphate
Capecitabine
Isonicotinic acid
DRUG METABOLISM - OTHER ENZYMES
6-Methyl-mercaptopurine
6-Methylthioinosine-5'-monophosphate
6-Methylthioguanosine monophosphate
6-Thioguanosine ...
|
|
hsa01040
|

|
Biosynthesis of unsaturated fatty acids
|
|
C16533 ((13Z,16Z)-Docosadienoic acid)
C16645 ((13Z,16Z)-Docosadienoyl-CoA)
C16645 ((13Z,16Z)-Docosadienoyl-CoA)
C03242 (Dihomo-gamma-linolenate)
C01595 (Linoleate)
C01530 (Octadecanoic acid)
C00712 ((9Z)-Octadecenoic ...
|
Docosadienoic acid
Δ13,16
Δ13,16
3.1.2.-
YciA
TesB
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.-
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2.2
3.1.2 ...
|
|
hsa01100
|

|
Metabolic pathways
|
|
C02647 (4-Guanidinobutanal)
C16848 (6-Deoxy-5-ketofructose 1-phosphate)
C22834 (Protein-trisulfide)
C16583 ((R)-(Homo)2-citrate)
C20581 (cis-(Homo)2-aconitate)
C06377 (D-Galactosamine 6-phosphate)
C20836 ...
|
Monoterpenoid biosynthesis
One carbon pool by folate
Flavonoid biosynthesis
Starch and sucrose metabolism
Biosynthesis of enediyne antibiotics
Toluene degradation
Tyrosine metabolism
Acarbose and validamycin ...
|
|
hsa01200
|

|
Carbon metabolism
|
Carbon metabolism is the most basic aspect of life. This map presents an overall view of central carbon metabolism, where the number of carbons is shown for each compound denoted by a circle, excluding ...
|
C00085 (D-Fructose 6-phosphate)
C01182 (D-Ribulose 1,5-bisphosphate)
C00197 (3-Phospho-D-glycerate)
C00101 (Tetrahydrofolate)
C00267 (alpha-D-Glucose), C00221 (beta-D-Glucose)
C00668 (alpha-D-Glucose 6-phosphate) ...
|
HCO
HCHO
HCHO
HCO
HCO
AcCoA
CARBON METABOLISM
Fructose-6P
Ribulose-1,5P
Glycerate-3P
THF
Glucose
Glucose-6P
Gluconate-6P
Fructose-1,6P
Glucono-1,5-lactone-6P
Glycerate-1,3P
Glyceraldehyde-3P
Glycerone-P
Glycerate-2P
Erythrose-4P
Sedohept-ulose-1 ...
|
|
hsa01210
|

|
2-Oxocarboxylic acid metabolism
|
2-Oxocarboxylic acids, also called 2-oxo acids and alpha-keto acids, are the most elementary set of metabolites that includes pyruvate (2-oxopropanoate), 2-oxobutanoate, oxaloacetate (2-oxosuccinate) and ...
|
C17254 (8-Methylthiooctyl glucosinolate)
C17232 (2-Oxo-10-methylthiodecanoic acid)
C17252 (7-Methylthioheptyl glucosinolate)
C17228 (2-Oxo-9-methylthiononanoic acid)
C17250 (Glucolesquerellin)
C17224 (2-Oxo-8-methylthiooctanoic ...
|
8-Methylthiooctyl glucosinolate
2-Oxo-10-methylthio-decanoic acid
7-Methylthioheptyl glucosinolate
2-Oxo-9-methylthio-nonanoic acid
6-Methylthiohexyl glucosinolate
2-Oxo-8-methylthio-octanoic acid
5-Methylthiopentyl ...
|
|
hsa01212
|

|
Fatty acid metabolism
|
|
C00024 (Acetyl-CoA)
C03939 (Acetyl-[acyl-carrier protein])
C05744 (Acetoacetyl-[acp])
C04618 ((3R)-3-Hydroxybutanoyl-[acyl-carrier protein])
C00083 (Malonyl-CoA)
C01209 (Malonyl-[acyl-carrier protein])
C04246 ...
|
FATTY ACID METABOLISM
Acetyl-CoA
Acetyl-[acp]
Malonyl-CoA
Malonyl-[acp]
Butanoyl-[acp]
(AcACP)
(MalACP)
MalACP
in mitochondria
Hexanoyl-[acp]
RM021
RM018 / RM020
MalACP
Octanoyl-[acp]
MalACP
Decanoyl-[acp]
MalACP
Dodecanoyl-[acp]
MalACP
Tetradecanoyl-[acp]
MalACP
Hexadecanoyl-[acp]
Palmitic ...
|
|
hsa01230
|

|
Biosynthesis of amino acids
|
This map presents a modular architecture of the biosynthesis pathways of twenty amino acids, which may be viewed as consisting of the core part and its extensions. The core part is the KEGG module for ...
|
C04390 (N6-Acetyl-LL-2,6-diaminoheptanedioate)
C04002 ((Z)-But-1-ene-1,2,4-tricarboxylate)
C00118 (D-Glyceraldehyde 3-phosphate)
C00111 (Glycerone phosphate)
C00236 (3-Phospho-D-glyceroyl phosphate)
C00197 ...
|
BIOSYNTHESIS OF AMINO ACIDS
Glyceraldehyde-3P
Glycerone-P
Glycerate-3P
Phosphoenolpyruvate
Pyruvate
Fructose-6P
Ribulose-5P
Ribose-5P
PRPP
Erythrose-4P
2-Oxo-butanoate
2-Oxoisovalerate
Valine
3-Methyl-2-oxopentanoate
Isoleucine
2-Oxoisocaproate
Leucine
Threonine
Serine
Glycine
Phosphoserine
O-Acetyl-serine
Cysteine
Aspartate
Homoserine
O-Succinylhomoserine
Cystathionine
Homocysteine
Methionine
Oxaloacetate
2 ...
|
|
hsa01232
|

|
Nucleotide metabolism
|
|
C00130 (IMP)
C00104 (IDP)
C00081 (ITP)
C00020 (AMP)
C00008 (ADP)
C00002 (ATP)
C00294 (Inosine)
C00212 (Adenosine)
C00147 (Adenine)
C00262 (Hypoxanthine)
C00242 (Guanine)
C00035 (GDP)
C00144 (GMP)
C00387 ...
|
IMP
IDP
ITP
AMP
ADP
ATP
Inosine
Adenosine
Adenine
Hypoxanthine
Guanine
GDP
GMP
Guanosine
GTP
Xanthine
Xanthosine
XMP
XTP
XDP
Deoxyadenosine
dAMP
dADP
dATP
dITP
dIDP
dIMP
Deoxyinosine
Deoxyguanosine
dGMP
dGTP
dGDP
Adenylo-succinate
degradation
UMP
UDP
UTP
Uridine
Uracil
CMP
CDP
CTP
Cytidine
Cytosine
dCMP
dCDP
dCTP
Deoxycytidine
dUMP
dUDP
dUTP
Deoxyuridine
dTMP
dTDP
dTTP
Thymidine
Thymine
degradation
degradation
NUCLEOTIDE ...
|
|
hsa01240
|

|
Biosynthesis of cofactors
|
|
C00118 (D-Glyceraldehyde 3-phosphate)
C00197 (3-Phospho-D-glycerate)
C00074 (Phosphoenolpyruvate)
C00022 (Pyruvate)
C00085 (D-Fructose 6-phosphate)
C00117 (D-Ribose 5-phosphate)
C00199 (D-Ribulose 5-phosphate)
C00119 ...
|
BIOSYNTHESIS OF COFACTORS
Glycer-aldehyde-3P
Glycerate-3P
Phosphoenol-pyruvate
Pyruvate
Fructose-6P
Ribose-5P
Ribulose-5P
PRPP
Serine
Cysteine
Aspartate
Cystathionine
Methionine
Oxaloacetate
Citrate
Isocitrate
2-Oxoglutarate
2-Oxoadipate
Glutamate
Glutamine
3-Dehydro-shikimate
Shikimate
Chorismate
Prephenate
4-Hydroxy-phenylpyruvate
Tyrosine
Tryptophan
AIR
IMP
GTP
Riboflavin
FMN
FAD
Ribulose-5P
Nicotinamide
Nicotinate
Nicotinate ...
|
|
hsa01250
|

|
Biosynthesis of nucleotide sugars
|
|
C04501 (N-Acetyl-alpha-D-glucosamine 1-phosphate)
C01170 (UDP-N-acetyl-D-mannosamine)
C06240 (UDP-N-acetyl-D-mannosaminouronate)
C20768 (UDP-2-acetamido-2,6-dideoxy-beta-L-talose)
C20774 (UDP-N-acetyl-beta-L-fucosamine)
C04613 ...
|
BIOSYNTHESIS OF NUCLEOTIDE SUGARS
GlcNAc-1P
UDP-ManNAc
UDP-ManNAcA
UDP-L-6dTalNAc
UDP-L-FucNAc
UDP-4-keto-6-deoxy-GlcNAc
UDP-D-FucNAc
UDP-D-QuiNAc
UDP-L-RhaNAc
UDP-L-QuiNAc
UDP-6-deoxy-AltNAc4N
UDP-6-deoxy-AltNAc4NAc
6-Deoxy-AltNAc4NAc
Pse5Ac7Ac
CMP-Pse5Ac7Ac
Neu5Ac
CMP-Neu5Ac
ManNAc
UDP-6-deoxy-L-IdoNAc4NAc
UDP-6-deoxy-L-GulNAc4NAc
6-Deoxy-L-GulNAc4NAc
8eLeg5Ac7Ac
CMP-8eLeg5Ac7Ac
UDP-GlcNAcA
UDP-3-keto-GlcNAcA
UDP-GlcNAc3NA
UDP-GlcNAc3NAcA
UDP-ManNAc3NAcA
UDP-ManNAc3NAmA
UDP-GalNAcA
UDP-GalNAc
UDP-GalNAcAN
dTDP-D-Glc
dTDP-4-oxo-6-deoxy-D-Glc
dTDP-4-oxo-6-deoxy-L-Man
dTDP-L-Rha
dTDP-6d-L-Tal
dTDP-3-oxo-6-deoxy-D-Gal
dTDP-Fuc3N
dTDP-Fuc3NAc
dTDP-Qui3N
dTDP-Qui3NAc
dTDP-3-oxo-6-deoxy-D-Glc
dTDP-Fuc4N
dTDP-Fuc4NAc
dTDP-4-oxo-6-deoxy-D-All
dTDP-6d-D-All
UDP-Gal
UDP-Galf
UDP-GlcA
CDP-Glc
CDP-4-keto-3 ...
|
|
hsa01521
|

|
EGFR tyrosine kinase inhibitor resistance
|
EGFR is a tyrosine kinase that participates in the regulation of cellular homeostasis. EGFR also serves as a stimulus for cancer growth. EGFR gene mutations and protein overexpression, both of which activate ...
|
C00165 (Diacylglycerol)
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
D07907 (Erlotinib (INN))
D01977 (Gefitinib (JP18/USAN/INN))
2065 (ERBB3), H00408 (Type 1 diabetes mellitus), H00865 (Lethal congenital ...
|
Non-small cell lung cancer
Her3
Her2
EGFR
Pancreatic cancer
mTOR signaling pathway
GSK-3
eIF-4EBP
p70S6K
mTOR
Her2
Bad
GAB1
TGFα
Her3
Sos
Grb2
Shc
PKC
PLCγ
PIP
PI3K
PKB/Akt
EGFR
MAPK signaling ...
|
|
hsa01522
|

|
Endocrine resistance
|
Endocrine therapy is a key treatment strategy to control or eradicate hormone-responsive breast cancer. The most commonly used endocrine therapy agents are selective estrogen receptor modulators (SERMs ...
|
C00951 (Estradiol-17beta)
C00951 (Estradiol-17beta)
C00575 (3',5'-Cyclic AMP)
C00951 (Estradiol-17beta)
C00951 (Estradiol-17beta)
D08559 (Tamoxifen (INN))
D00964 (Letrozole (JAN/USP/INN)), D00960 (Anastrozole ...
|
ENDOCRINE RESISTANCE
GPR30
Src
MMP
HB-EGF
EGFR
HB-EGF
CoA
DNA
DNA
cAMP
PKA
Ras
Raf
MEK
ERK1/2
GPR30
MAPK signaling pathway
Shc
Grb2
SOS
Gβγ
Loss or modification of ER
Ovarian steroidogenesis
PI3K
Akt
Alternative ...
|
|
hsa01523
|

|
Antifolate resistance
|
Since the 1940s, antifolates have played a pivotal role in drug treatment of malignant, microbial, parasitic and chronic inflammatory diseases. The molecular basis of the anti-proliferative activity of ...
|
D00142 (Methotrexate (JP18/USP/INN)), D05589 (Pralatrexate (JAN/USAN/INN)), D04766 (Lometrexol sodium (USAN)), D01064 (Raltitrexed (JAN/USAN/INN)), D07472 (Pemetrexed (INN)), D05400 (Pelitrexol (USAN/INN)) ...
|
ANTIFOLATE RESISTANCE
RFC
ABCC1
ABCC2
ABCC3
ABCC4
ABCC5
GARTF
Antifolate
Methotrexate (MTX)
Pemetrexed (MTA)
Pralatrexate (PDX)
Lometrexol (LMX)
AG2034
GW1843
BGC945
Trimetrexate
Endosome
Antifolate
Decreased ...
|
|
hsa01524
|

|
Platinum drug resistance
|
Platinum-based drugs cisplatin, carboplatin and oxaliplatin are widely used in the therapy of solid malignancies, including testicular, ovarian, head and neck, colorectal, bladder and lung cancers. The ...
|
D00275 (Cisplatin (JP18/USP/INN))
D01790 (Oxaliplatin (JAN/USP/INN))
D01363 (Carboplatin (JP18/USP/INN))
472 (ATM), H00005 (Chronic lymphocytic leukemia), H00064 (Ataxia telangiectasia), H00094 (Immunodeficiency ...
|
Mitochondrion
p53 signaling pathway
ATM
p53
Bcl-2
CASP9
CASP3
Bad
IAP/XIAP
Apaf-1
CytC
Bid
CASP8
FADD
Fas
PLATINUM DRUG RESISTANCE
Fas-L
DNA
Pro-apoptotic genes
DNA Damage
Apoptosome
Bax
Bak
NOXA
PUMA
Fas
tBid
Bax
Bak
Bax/Bak-induced ...
|
|
hsa02010
|

|
ABC transporters
|
The ATP-binding cassette (ABC) transporters form one of the largest known protein families, and are widespread in bacteria, archaea, and eukaryotes. They couple ATP hydrolysis to active transport of a ...
|
C16421 (AI-2)
C01684 (D-Rhamnose)
C00095 (D-Fructose)
C01487 (D-Allose)
C00181 (D-Xylose)
C03619 (Methyl beta-D-galactoside)
C00259 (L-Arabinose)
C00140 (N-Acetyl-D-glucosamine)
C00185 (Cellobiose)
C05402 ...
|
GguA
GguB
ChvE
LsrA
LsrD
LsrC
LsrB
Autoinducer 2
RhaT
RhaQ
RhaP
RhaS
Rhamnose
FrcA
FrcC
FrcB
Fructose
AlsA
AlsC
AlsB
D-Allose
XylG
XylH
XylF
D-Xylose
MglA
MglC
MglB
Methyl-galactoside
AraG
AraH
AraF
L-Arabinose
N-Acetylglucosamine
Cellobiose
Raffinose/ ...
|
|
hsa03008
|

|
Ribosome biogenesis in eukaryotes
|
Ribosomes are the cellular factories responsible for making proteins. In eukaryotes, ribosome biogenesis involves the production and correct assembly of four rRNAs and about 80 ribosomal proteins. It requires ...
|
100008588 (RNA18SN5), 106631781 (RNA18SN1), 109864271 (RNA45SN4), 109864273 (RNA18SN4), 109864280 (RNA18SN2), 109910380 (RNA18SN3)
100008587 (RNA5-8SN5), 106632260 (RNA5-8SN1), 109864274 (RNA5-8SN4), 109864281 ...
|
RIBOSOME BIOGENESIS IN EUKARYOTES
Pol I
Pol III
Nucleolus
UTP-B complex
18S
5.8S
25S
U3 complex
MPP10
Imp4
Imp3
UTP9
NAN1
UTP15
UTP4
UTP10
UTP8
UTP5
UTP18
UTP13
UTP6
PWP2
UTP21
Dip2
CK2A
CK2B
UTP22
Rrp7
t-UTP ...
|
|
hsa03010
|

|
Ribosome
|
|
2197 (FAU)
6137 (RPL13), H02187 (Spondyloepimetaphyseal dysplasia)
6158 (RPL28)
6208 (RPS14), H01484 (5q- syndrome)
6228 (RPS23), H02637 (Brachycephaly, trichomegaly, and developmental delay)
6222 (RPS18)
6235 ...
|
L7A
S30e
L13e
L28e
S14e
S23e
S18e
S29e
S13e
S11e
S15e
S6e
S8e
S17e
S19e
S24e
S27e
S27Ae
S7e
S10e
S12e
S21e
S25e
S26e
S28e
S4e
S3Ae
25S
23S
L10
L11
L3e
L11e
L9e
LP0
L12e
L13Ae
L13
5.8S
L10Ae
L8e
L14
L23e
L27Ae
L15
L5e
L18
L17e
L22
L23Ae
L23
L26e
L24
L35e
L29
L30
L7e
L15e
L18e
L19e
L21e
L24e
L30e
L39e
L40e
L41e
L44e
L31e
L32e
L34e
L35Ae
L37e
L37Ae
L6e
L18Ae
L22e
L27e
L29e
L36e
L38e
L4e
L7Ae
L10e
L14e
L7/L12
L12
LP1 ...
|
|
hsa03013
|

|
Nucleocytoplasmic transport
|
The exchange of molecules between the nucleus and cytoplasm is mediated through nuclear pore complexes (NPCs) embedded in the nuclear envelope. The NPC is composed of highly conserved, distinct structural ...
|
11260 (XPOT), 1434 (CSE1L), 23039 (XPO7), 23214 (XPO6), 57510 (XPO5), 64328 (XPO4), 64901 (RANBP17), 7514 (XPO1), D11222 (Selinexor (USAN/INN)), D11223 (Verdinexor (USAN/INN)), D11499 (Eltanexor (USAN/INN))
5901 ...
|
NES
NUCLEOCYTOPLASMIC TRANSPORT
Ran GTP
XPO
Ran
CBC
PHAX
Central channel
Nuclear basket
Cytoplasmic ring
Cytoplasmic fibrils
Spoke complex
Lumenal ring
Nucleoplasmic ring
Central channel
Nup58/45
Nup205
Nup188
Nup62
Tpr
Nup153
Nup93
Nup54
Spoke ...
|
|
hsa03015
|

|
mRNA surveillance pathway
|
The mRNA surveillance pathway is a quality control mechanism that detects and degrades abnormal mRNAs. These pathways include nonsense-mediated mRNA decay (NMD), nonstop mRNA decay (NSD), and no-go decay ...
|
5976 (UPF1)
26019 (UPF2)
65109 (UPF3B), 65110 (UPF3A), H00658 (X-linked syndromic intellectual developmental disorder)
23049 (SMG1)
23381 (SMG5)
9887 (SMG7)
23293 (SMG6)
2107 (ETF1)
23708 (GSPT2), 2935 ...
|
Spliceosome
m7G
m7G
EJC complex
m7G
Upf3
Tap
40S
SRm160
Pinin
Ref/Aly
Tap
Upf1
Upf2
Upf3
SMG1
SMG5
SMG7
SMG6
Ribosome
eRF1
eRF1
eRF3
CBC
Splicing
mRNA SURVEILLANCE PATHWAY
Recognition of PTC (premature ...
|
|
hsa03018
|

|
RNA degradation
|
The correct processing, quality control and turnover of cellular RNA molecules are critical to many aspects in the expression of genetic information. In eukaryotes, two major pathways of mRNA decay exist ...
|
2023 (ENO1), 2026 (ENO2), 2027 (ENO3), H00069 (Glycogen storage disease), H01762 (Muscle glycogen storage disease), H01953 (Glycogen storage disease type XIII)
87178 (PNPT1), H00605 (Deafness, autosomal ...
|
RNase E
RNase E
RNase E
RhlB
Enolase
PNPase
RhlE
RNaseR
helicases
Rho
RNA degradosome type A
(Escherichia coli)
RNA degradosome type B
(Pseudomonas)
RNA degradosome type C
(Rhodobacter)
Csl4 / Rrp4
Csl4 ...
|
|
hsa03020
|
|
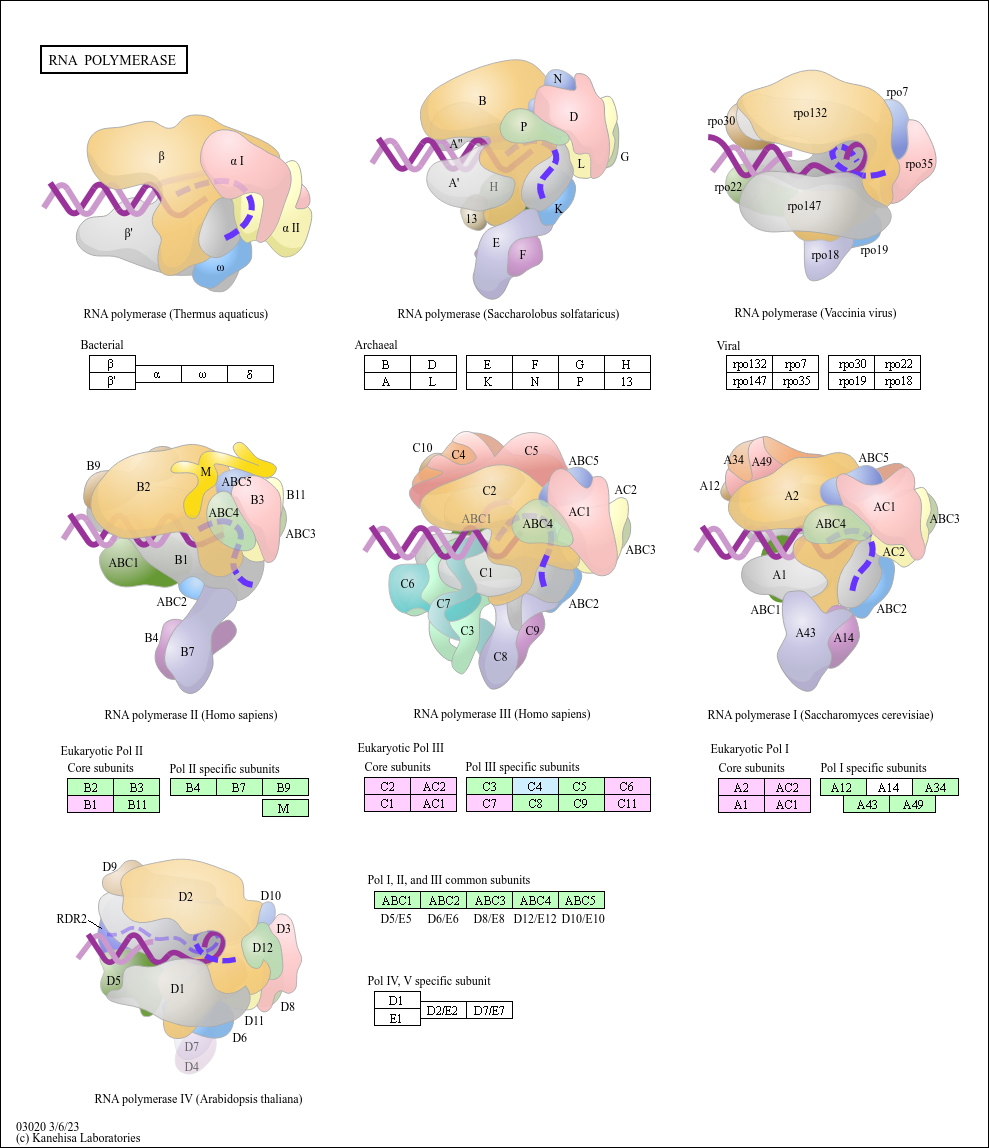
RNA polymerase
|
|
5431 (POLR2B)
5430 (POLR2A), H02571 (Neurodevelopmental disorder with hypotonia and variable intellectual and behavioral abnormalities)
5432 (POLR2C)
5433 (POLR2D)
5434 (POLR2E)
5435 (POLR2F)
5436 (POLR2G)
5437 ...
|
RNA POLYMERASE
ABC1
ABC2
ABC3
ABC5
B11
ABC4
RNA polymerase (Thermus aquaticus)
RNA polymerase II (Homo sapiens)
Bacterial
Eukaryotic Pol II
α I
α II
ABC3
ABC1
B11
ABC4
ABC5
Core subunits
Pol I II, and ...
|
|
hsa03022
|

|
Basal transcription factors
|
|
387332 (TBPL2), 6908 (TBP), 9519 (TBPL1), H00063 (Spinocerebellar ataxia (SCA)), H01243 (Huntington disease-like syndrome)
138474 (TAF1L), 6872 (TAF1), H00658 (X-linked syndromic intellectual developmental ...
|
General transcription factors for RNA polymerase II
BASAL TRANSCRIPTION FACTORS
TBP
TAF1
TAF2
TAF3
TAF4
TAF5
TAF6
TAF7
TAF8
TAF9
TAF10
TAF11
TFIIB
TFIIA1
TFIIA2
TFII-I
TFIIF1
TFIIF2
XPD
TFIIE1
TFIIE2
TFIIH1
TFIIH2
TFIIH3
TFIIH4
TATA
Pol ...
|
|
hsa03030
|
|
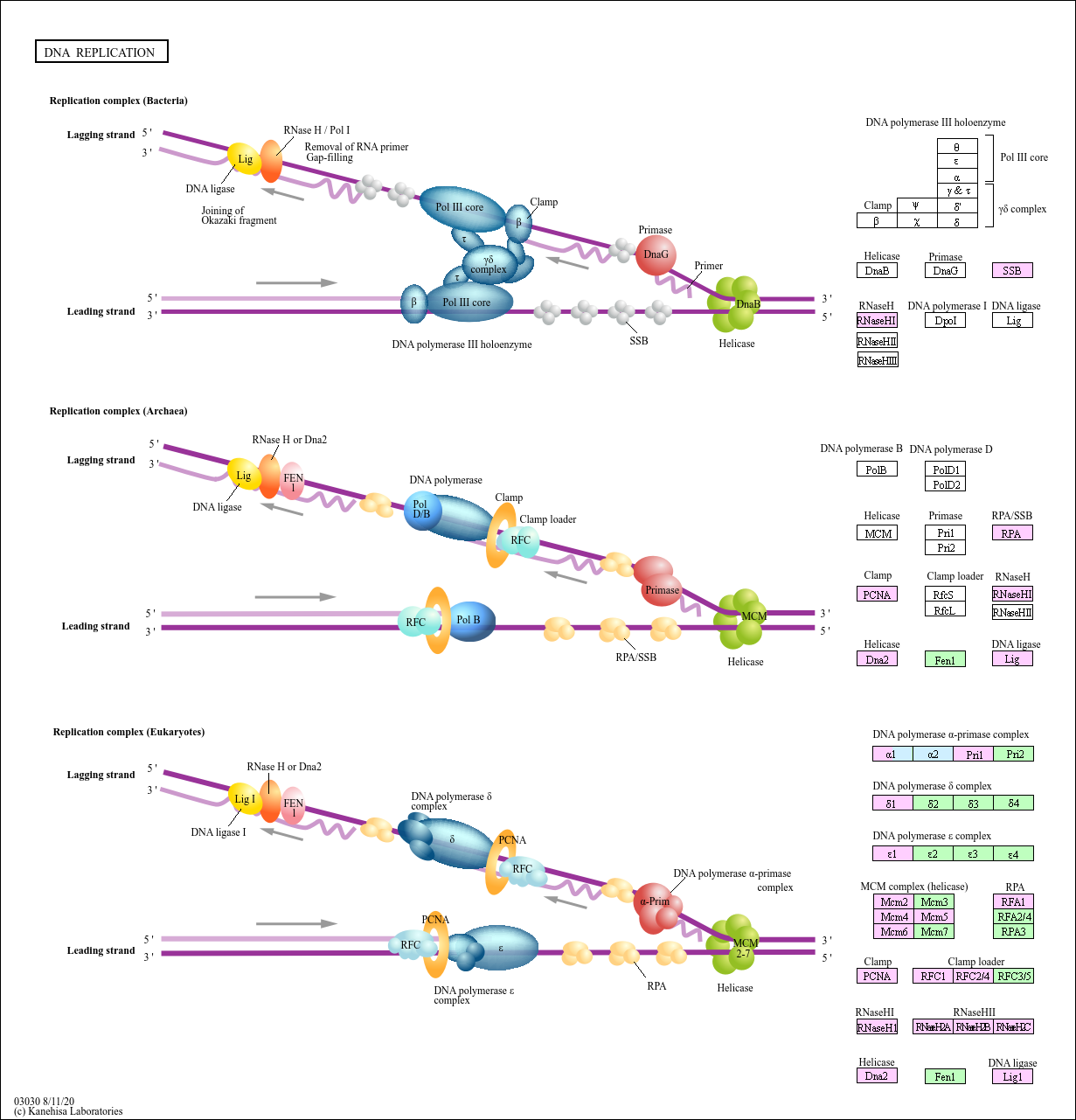
DNA replication
|
A complex network of interacting proteins and enzymes is required for DNA replication. Generally, DNA replication follows a multistep enzymatic pathway. At the DNA replication fork, a DNA helicase (DnaB ...
|
3978 (LIG1), H00094 (Immunodeficiency associated with DNA repair defects)
2237 (FEN1)
1763 (DNA2), H00992 (Seckel syndrome), H01118 (Progressive external ophthalmoplegia), H01734 (Rothmund-Thomson syndrome)
10535 ...
|
Lig1
Fen1
Dna2
RNaseH2A
RNaseH2B
RFC2/4
RNaseH2C
RFC3/5
PCNA
RPA3
Mcm5
SSB
DnaG
DNA REPLICATION
γ & τ
RNaseHI
Lig
Mcm2
Mcm3
Mcm4
Pri1
Pri2
Mcm6
Mcm7
RFA1
RFA2/4
RFC1
DpoI
DnaB
Replication complex ...
|
|
hsa03040
|

|
Spliceosome
|
After transcription, eukaryotic mRNA precursors contain protein-coding exons and noncoding introns. In the following splicing, introns are excised and exons are joined by a macromolecular complex, the ...
|
9879 (DDX46)
7919 (DDX39B)
8449 (DHX16), H02745 (Neuromuscular oculoauditory syndrome), H02747 (Oculogastrointestinal neurodevelopmental syndrome)
1659 (DHX8)
1665 (DHX15)
9785 (DHX38), H00527 (Retinitis ...
|
complex
pre-mRNA
Exon
5' splice site
3' splice site
Branch point
complex
complex
complex B*
Post-spliceosomal complex
mRNA
SPLICEOSOME
U2AF
Activated spliceosome
PRP19 complex
U4/U6.U5 tri-snRNP
Intron
Prp5
UAP56
Prp2
Prp22
Prp43
...
|
|
hsa03050
|

|
Proteasome
|
The proteasome is a protein-destroying apparatus involved in many essential cellular functions, such as regulation of cell cycle, cell differentiation, signal transduction pathways, antigen processing ...
|
3458 (IFNG), H00083 (Allograft rejection), H00084 (Graft-versus-host disease), H00342 (Tuberculosis), H01132 (Aplastic anemia), H01563 (HIV infection), D04242 (Fontolizumab (USAN/INN)), D11120 (Emapalumab ...
|
IFNγ
Rpn1
Rpn2
Rpn10
Rpn11
Lid
Base
20S
Core Particle
Regulatory Particle
19S
26S proteasome
(PA700)
(PA700-20S-PA700)
Regulatory Particles
Rpn3
Rpn5
Rpn6
Rpn7
Rpn8
Rpn9
Rpn11
Rpn12
PA700 (Lid)
Rpn15
Rpn10
Rpn1
Rpn2
Rpn13
Rpt1
Rpt2
Rpt3
Rpt4
Rpt5
PA700 ...
|
|
hsa03060
|

|
Protein export
|
The protein export is the active transport of proteins from the cytoplasm to the exterior of the cell, or to the periplasmic compartment in Gram-negative bacteria. The sec dependent pathway is the general ...
|
5018 (OXA1L)
6727 (SRP14)
6731 (SRP72), H02529 (Bone marrow failure syndrome)
6730 (SRP68), H00100 (Neutropenic disorders)
6729 (SRP54), H00100 (Neutropenic disorders)
6728 (SRP19)
6734 (SRPRA)
6726 (SRP9)
6729 ...
|
TatE
Ffs
YidC
TatB
TatC
TatA
SecA
SecY
SecE
SecG
SecD/F
YajC
SPase I
SPase II
SRP14
SRP72
SRP68
SRP54
SRP19
SRPR
SRP9
SecB
PROTEIN EXPORT
Sec dependent pathway
Signal peptidase
TAT (twin-arginine translocation) ...
|
|
hsa03082
|

|
ATP-dependent chromatin remodeling
|
|
57492 (ARID1B), 8289 (ARID1A), H00048 (Hepatocellular carcinoma), H00773 (Autosomal dominant intellectual developmental disorder), H01403 (Coffin-Siris syndrome)
8110 (DPF3), 8193 (DPF1)
5977 (DPF2), H01403 ...
|
ARID1
DPF1/3
DPF2
SS18
BAZ2B
BAZ2A
BAZ1A
BAZ1B
BAZ1A
ATP - DEPENDENT CHROMATIN REMODELING
Chromatin remodeling complexes (CRCs)
ISWI family
ACF
WICH
NORC
BRF
CHRAC
RSF
NURF
CERF
CHD family
NURD complex
SWI ...
|
|
hsa03083
|

|
Polycomb repressive complex
|
|
2145 (EZH1), D11551 (Valemetostat (INN)), D11662 (Valemetostat tosilate (JAN))
2146 (EZH2), H01613 (Follicular lymphoma), H01751 (Weaver syndrome), H02410 (Myelodysplastic/myeloproliferative neoplasms) ...
|
POLYCOMB REPRESSIVE COMPLEX
Crosstalk between cPRC1 and PRC2
PRC2
Crosstalk between ncPRC1 and PRC2
H2AK119ub-independent chromatin compaction
H3K27me3
H2AK119ub1
nucleosome
CpG
PRC2
EZH
cPRC1
CBX
RNF
Target ...
|
|
hsa03250
|

|
Viral life cycle - HIV-1
|
|
11168 (PSIP1)
7251 (TSG101)
128866 (CHMP4B), 29082 (CHMP4A), 79643 (CHMP6), H01202 (Cataract)
10015 (PDCD6IP), H00269 (Primary microcephaly)
1234 (CCR5), H00408 (Type 1 diabetes mellitus), H00413 (Hepatitis ...
|
Binding
Preintegration complex (PIC)
reverse transcriptase
matrix
integrase
Fusion
Reverse transcription
nuclear import
Integration
LTR
RT(RNase H)
Reverse transcriptase complex (RTC)
p51
tat
vpr
nef
vif
vpr
viral ...
|
|
hsa03260
|

|
Virion - Human immunodeficiency virus
|
|
C00925 (Heparan sulfate)
920 (CD4), H02526 (Disorders of adaptive immunity), D03420 (Cedelizumab (USAN/INN)), D05610 (Priliximab (USAN/INN)), D06356 (Zanolimumab (USAN/INN)), D09190 (Anti-human thymocyte ...
|
VIRION - HUMAN IMMUNODEFICIENCY VIRUS
matrix
capsid
nucleocapsid
gag
gagpol
Structural protein
tat
vpu
gp41
gp120
Polyprotein
Regulatory protein
Envelope glycoprotein
Receptor
CD4
CCR5
CXCR4
CLEC4M
CD209
hiv-1
rev
Viral ...
|
|
hsa03264
|

|
Virion - Flavivirus
|
|
C00925 (Heparan sulfate), C00607 (Chondroitin sulfate)
26762 (HAVCR1)
7301 (TYRO3), D12650 (Zanzalintinib (USAN/INN)), D12651 (Zanzalintinib fumarate (USAN))
10332 (CLEC4M), 30835 (CD209), H00342 (Tuberculosis) ...
|
NS3
NS4A
NS5
RdRP / MTase
NS2B
NS4B
NS1
NS2A
Zika virus
Nonstructural protein
Structural protein
HAVCR1
TYRO3
CD209
Receptor
POLY
Polyprotein
VIRION - FLAVIVIRUS
membrane
capsid
envelope
helicase ...
|
|
hsa03265
|
|
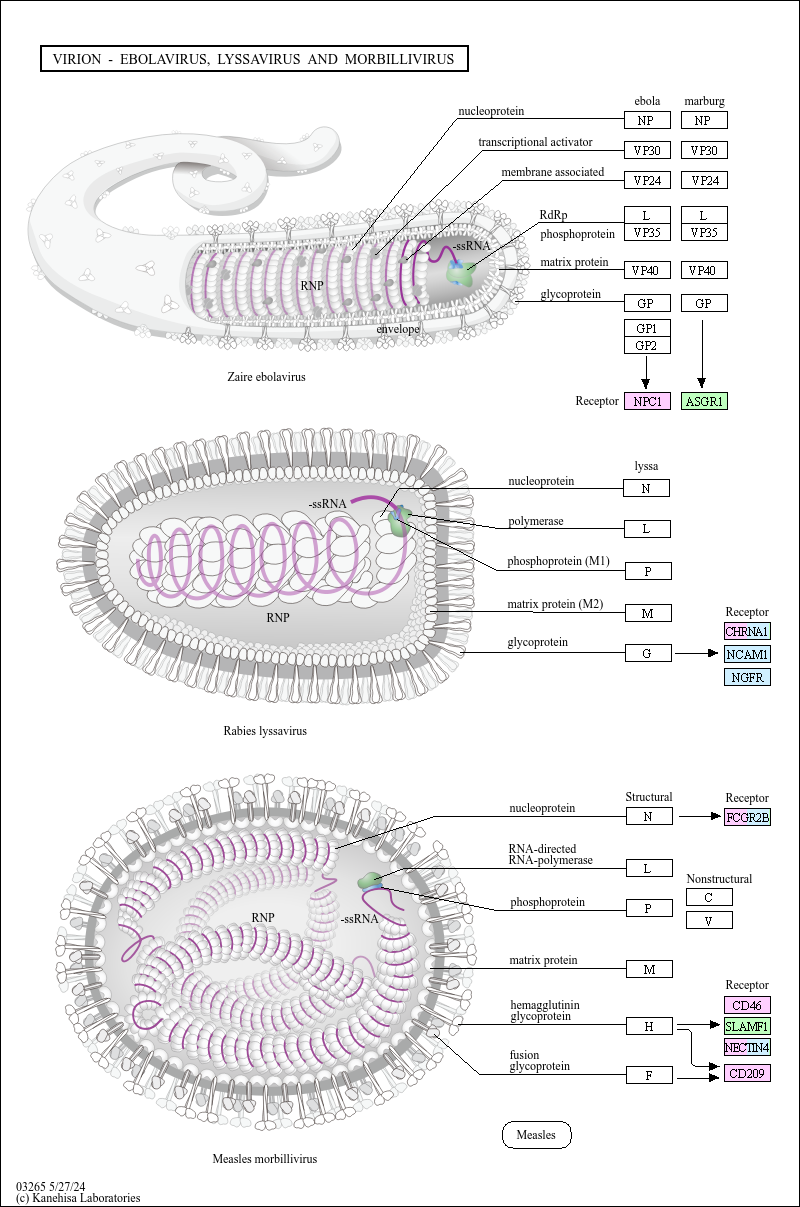
Virion - Ebolavirus, Lyssavirus and Morbillivirus
|
|
4804 (NGFR), D11028 (Cenegermin (INN))
1134 (CHRNA1), H00770 (Congenital myasthenic syndrome), H00986 (Multiple pterygium syndrome), D00611 (Mecamylamine hydrochloride (USP)), D00761 (Metocurine iodide ...
|
Rabies lyssavirus
VIRION - EBOLAVIRUS AND LYSSAVIRUS
NGFR
CHRNA1
NCAM1
Receptor
matrix protein (M2)
polymerase
phosphoprotein (M1)
nucleoprotein
glycoprotein
-ssRNA
ribonucleoprotein (RNP)
Zaire ebolavirus
-ssRNA
ribonucleoprotein ...
|
|
hsa03266
|

|
Virion - Herpesvirus
|
|
3678 (ITGA5), D06319 (Volociximab (USAN)), D12722 (Bexotegrast (USAN/INN))
3690 (ITGB3), H00083 (Allograft rejection), H00226 (Glanzmann thrombasthenia), H01235 (Bleeding disorder platelet-type), H01730 ...
|
VIRION - HERPESVIRUS
Receptor
Capside protein
UL27
UL1
VP5
VP19C
VP23
UL19
UL38
UL18
major capsid
triplex
CVSC
small capsid
UL17
UL25
UL35
UL6
portal vertex
UL36
UL37
UL7
UL51
Glycoprotein
UL22
UL10
UL49 ...
|
|
hsa03267
|

|
Virion - Adenovirus
|
|
1525 (CXADR)
941 (CD80), D03203 (Abatacept (USAN/INN)), D03222 (Belatacept (USAN/INN)), D04295 (Galiximab (USAN/INN)), D12463 (Davoceticept (USAN/INN))
942 (CD86), D03203 (Abatacept (USAN/INN)), D03222 ...
|
VIRION - ADENOVIRUS
Major capsid protein
Minor / Cement protein
Receptor
Fiber
CXADR
CD80
CD86
CD46
IIIa
VIII
Core protein
VII
IVa2
preterminal protein
PTP
52K
Human adenovirus
dsDNA
penton ...
|
|
hsa03271
|

|
Virion - Rotavirus
|
|
3673 (ITGA2), H01235 (Bleeding disorder platelet-type)
3688 (ITGB1), D06319 (Volociximab (USAN)), D09799 (Carotegrast methyl (JAN)), D10028 (Valategrast hydrochloride (USAN)), D12722 (Bexotegrast (USAN/INN))
|
VIRION - ROTAVIRUS
Rotavirus A
dsRNA
Structural
VP1
RdRp
VP3
cap
core
VP2
inner
VP6
outer
VP7
spike
VP4
VP5
VP8
ITGA2
ITGB1
Receptor
NSP1
Nonstructural
NSP2
NSP3
NSP4
NSP5
NSP6
SLP
DLP
TLP
capsid
|
|
hsa03320
|

|
PPAR signaling pathway
|
Peroxisome proliferator-activated receptors (PPARs) are nuclear hormone receptors that are activated by fatty acids and their derivatives. PPAR has three subtypes (PPARalpha, beta/delta, and gamma) showing ...
|
C02165 (Leukotriene B4), C14776 (8(S)-HETE)
C15493 (9-cis-Retinoic acid)
C15493 (9-cis-Retinoic acid)
C14767 (9(S)-HODE), C14762 (13(S)-HODE)
C15493 (9-cis-Retinoic acid)
100509620 (AQP7B), 364 (AQP7) ...
|
Unsaturated fatty acid
Unsaturated fatty acid
Saturated fatty acid
Eicosanoid
Fibrate drug
NSAID
9-cis-Retinoic acid
Unsaturated fatty acid
9-cis-Retinoic acid
NSAID
AQP7
GyK
PEPCK
Perilipin
FATCD36
FATP
FABP
PPARα
RXR
PPARβ/δ
RXR
PPARγ
RXR
SCP-X
Thiolase ...
|
|
hsa03410
|

|
Base excision repair
|
Base excision repair (BER) is the predominant DNA damage repair pathway for the processing of small base lesions, derived from oxidation and alkylation damages. BER is normally defined as DNA repair initiated ...
|
328 (APEX1)
2237 (FEN1)
3978 (LIG1), H00094 (Immunodeficiency associated with DNA repair defects)
3980 (LIG3), H00469 (Mitochondrial DNA depletion syndrome), H01390 (Mitochondrial neurogastrointestinal ...
|
APEX
Fen1
Lig1
Lig3
Lig
Polε
Polδ
Polβ
Polβ
Polβ
Polβ
PCNA
PCNA
PCNA
DpoI
DpoI
DpoI
APEX
Nfo
Xth
MUTY
UNG
NTH
Nei
Fpg
Fpg
Polβ
Lig
XRCC1
Lig3
HMGB1
Lig1
PCNA
Polλ
RecJ
Fen1
XRCC1
OGG1
BASE EXCISION REPAIR
MBD4
Tag
MPG
Mug
SMUG
NEIL
Xth
APEX
Nfo
Polδ/ε
PARP
AlkA
APEX
XRCC1
DpoI
Nei
Oxidized ...
|







































